Biosignal Processing for Human-Machine Interaction
Pre-shot learning techniques to read your mind and biosignals for calibration-free brain-computer interface (BCI) and human-machine interaction (HMI).
MERL Researchers:
Ye Wang,
Toshiaki Koike-Akino.
Joint work with: Deniz Erdogumus (Northeastern University)
Search MERL publications by keyword:
deep learning,
brain,
EEG,
EMG,
biosignal,
Mind Sensing for Human-Machine Interaction (HMI)
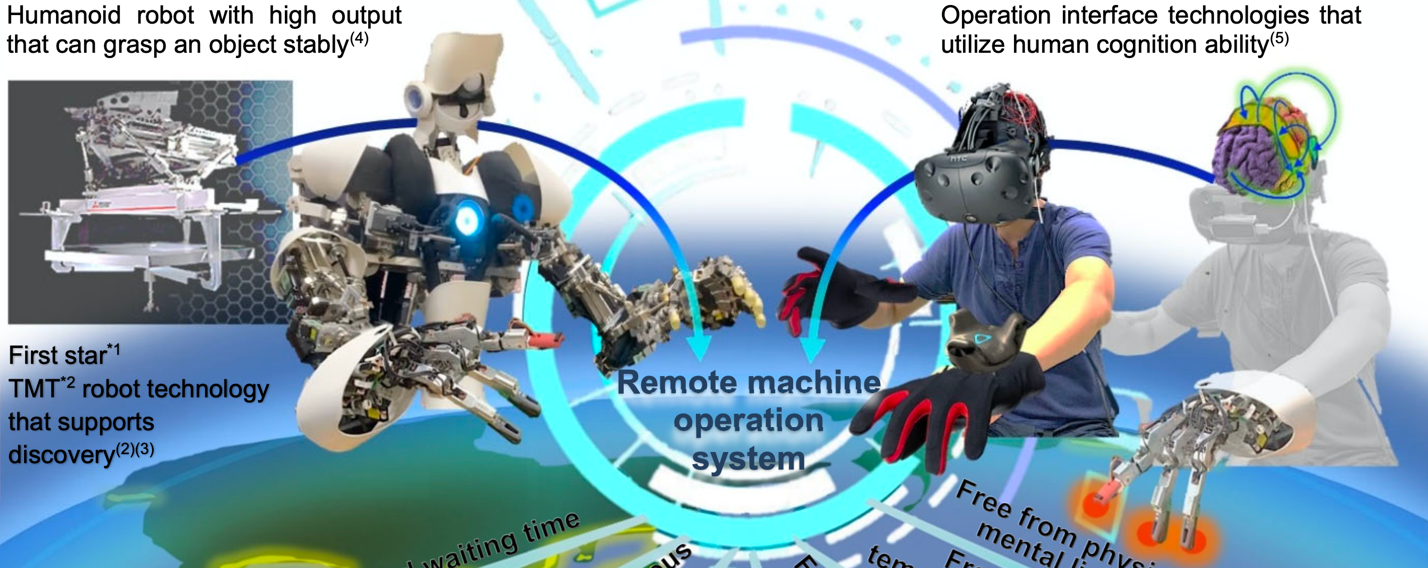
Realizing Sci-Fi scenes---an intelligent robot is able to read your thoughts in your mind---may be no longer a far future dream thanks to rapid progress in robotics, sensors, and artificial intelligence (AI). Biosignal processing to analyze human's physiological states is a key enabling technology for mind sensing in HMI and BCI systems. When machine intelligence can collaboratively support human intelligence without conflict, HMI systems will make a breakthrough in various scenarios, including teleworking, maintaining remote facilities, disaster response, epidemic care, exoskeletons, education, etc. MERL has been developing fundamental technologies in biosignal processing to analyze vital signs, physiological data, and biometrics to enable improved HMI systems.
Biometrics and Biosignal Processing
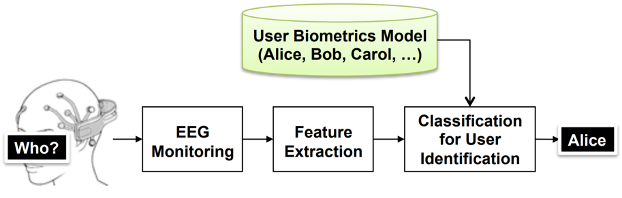
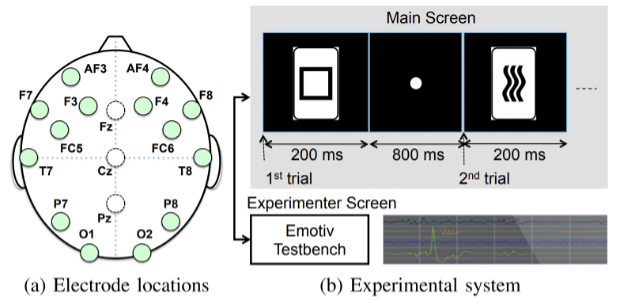
Biosignals are signals that can be measured or monitored in living beings including, but not limited to, electroencephalography (EEG), electrocorticography (ECoG), electrocardiography (ECG), and electromyography (EMG). In a similar way as human fingerprints, biosignals are sometimes used for user identification and authentication biometrics. MERL has demonstrated that EEG signal processing can accurately identify individuals (TR2016-105). Nonetheless, biosignals highly depend on the subject's physiological conditions, mental states, and various other factors. The subject dependency of biometrics is a major nuisance factor for developing universal HMI systems which can be seamlessly used across different users.
Transfer-Ready Pre-Shot Learning
To deal with subject-dependent nuisance factors, typical HMI systems require long machine-calibration sessions or long human-training sessions. Transfer learning (TL) and domain adaptation (DA) across different users are key AI techniques to facilitate the calibration/training sessions. Beyond typical few-shot learning and zero-shot learning used as the state-of-the-art TL methods, our development is specialized in a new paradigm called pre-shot learning (a.k.a., domain generalization), which does not require calibration for new users.
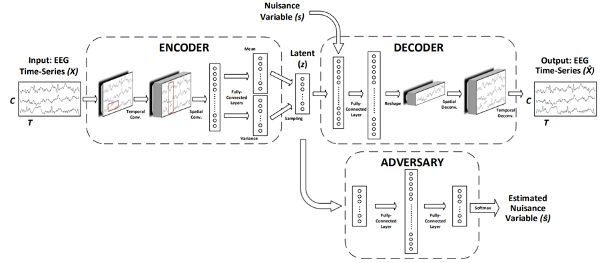
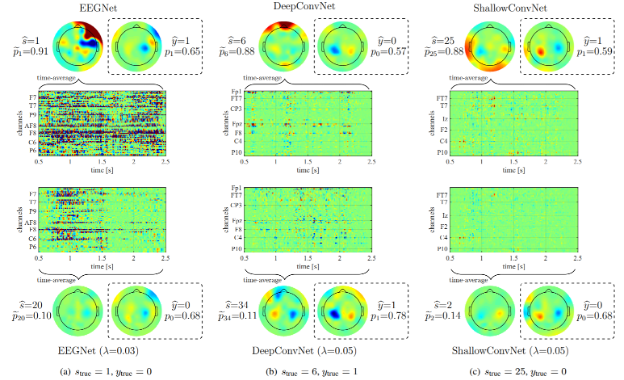
In pre-shot learning, nuisance-robust/invariant features are extracted from biosignals by disentangling subject-dependent factors via adversarial training and other regularization techniques. Our transfer-ready feature extraction for domain-invariant learning has been verified as effective for HMI development. Some of our recent achievements are listed below:
- Adversarial autoencoder to extract nuisance-invariant feature (TR2018-184)
- Adversarial censoring for transfer-ready classification across users and sessions (TR2019-017, TR2020-049)
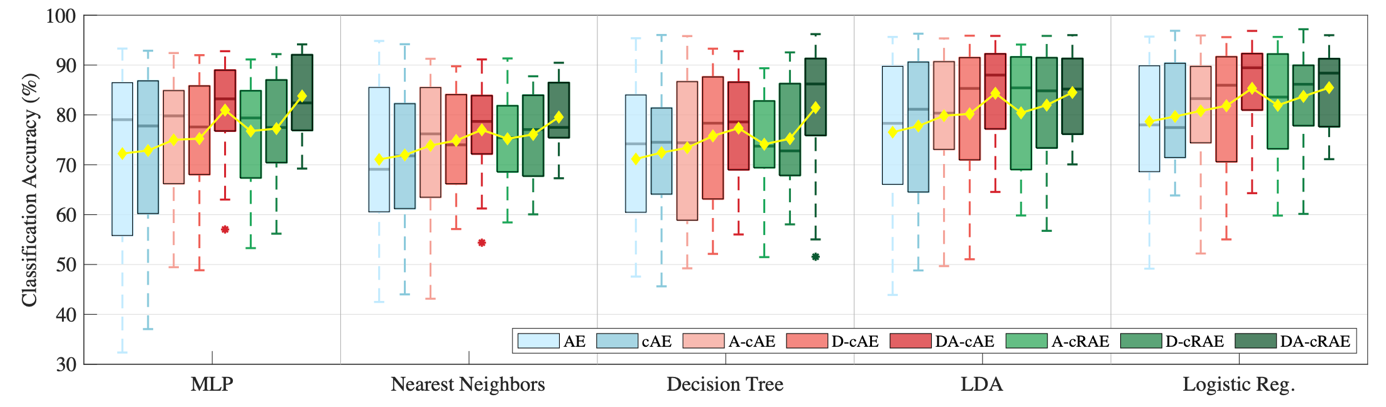
- Complementary adversarial entanglement to exploit both subject-invariant/dependent features (TR2020-109, TR-2020-128)
- Rateless adversarial censoring for hyperparameter-insensitive pre-shot learning (TR2021-027)
- Few-shot calibration to ensemble user-specific models (TR2018-210)
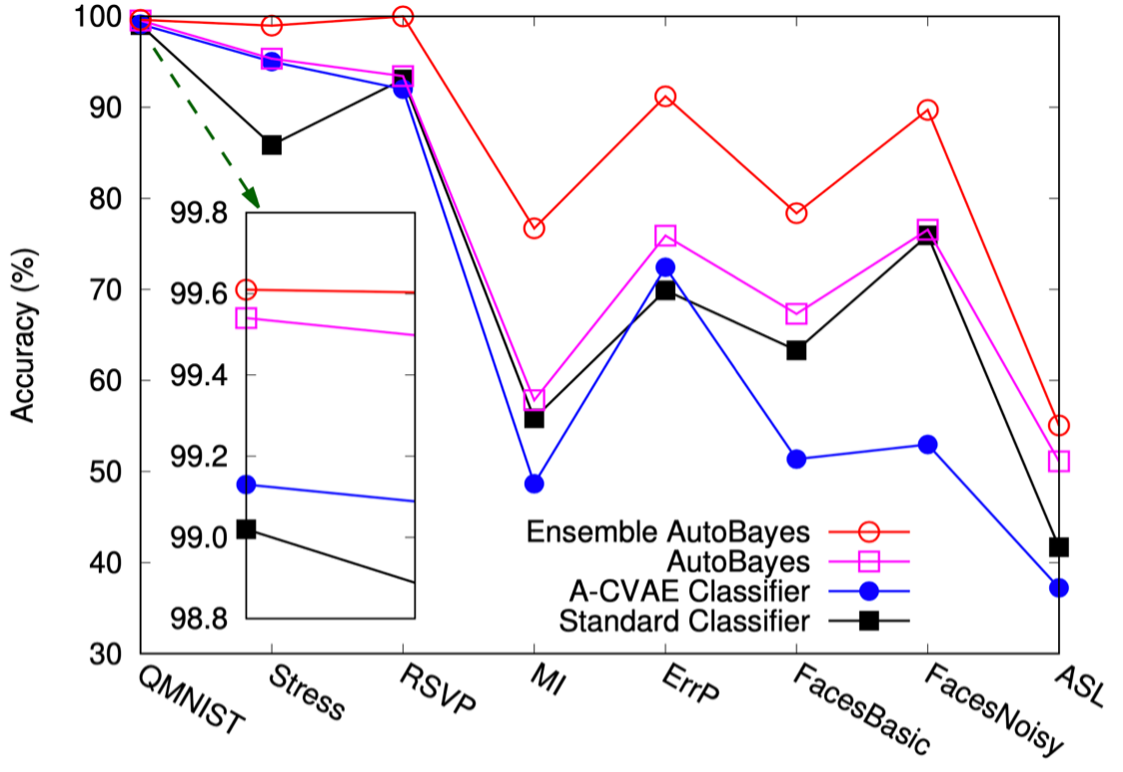
- Justifiable automated adversarial Bayesian inference: AutoBayes (TR2021-016)
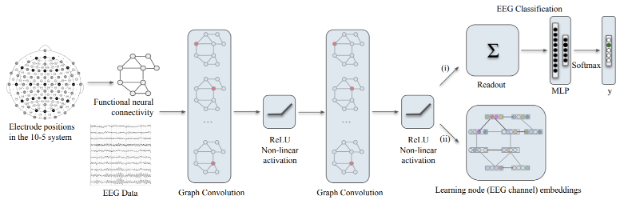
- Graph neural network (GNN) inspired by cognitive geometry (TR2021-PENDING)
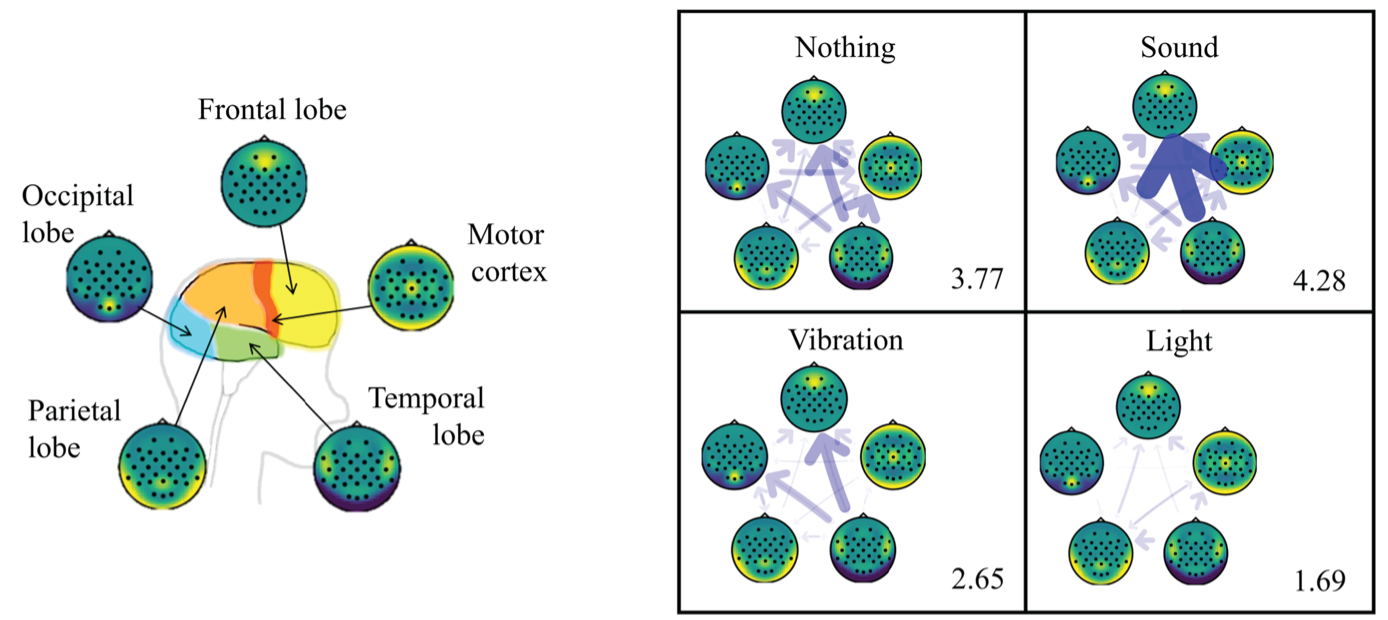
- Cognitive information flow for visual haptics interface (TR2021-031, TR2021-063)
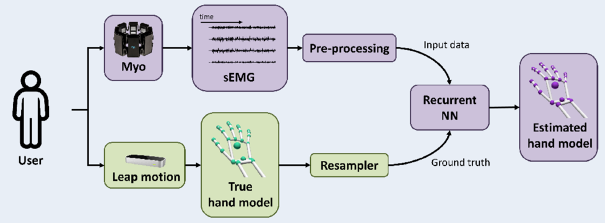
- Probabilistic EMG motion tracking for bionic hands control (TR2018-014)
- User identification and authentication with EEG biometrics (TR2016-105)
- Automated subject transfer learning: AutoTransfer (Top score in cross-subject transfer task at NeurIPS competition)
Videos
Media Coverage
- Nikkan Kogyou Shimbun (Japanese): https://newswitch.jp/p/24993
- Dempa Shimbun (Japanese): https://dempa-digital.com/article/151793
Related Publications
- Invariant Representations from Adversarially Censored Autoencoders:
https://arxiv.org/abs/1805.08097 - Proposal and Evaluation of Visual Haptics for Manipulation of Remote Machine System:
https://www.frontiersin.org/articles/10.3389/frobt.2020.529040/full - Remote Machine System with Sense of Oneness:
https://www.giho.mitsubishielectric.co.jp/giho/pdf/2021/2104106.pdf - Human x Machine Remote Fusion System: Visual Haptics Implementation:
https://robogaku.jp/news/2021/rsj2020_1b1.html - Remote Machine System with Sense of Oneness:
http://www.interaction-ipsj.org/proceedings/2020/data/pdf/1P-74.pdf
MERL News & Events
-
AWARD MERL Ranked 1st Place in Cross-Subject Transfer Learning Task and 4th Place Overall at the NeurIPS2021 BEETL Competition for EEG Transfer Learning. Date: November 11, 2021
Awarded to: Niklas Smedemark-Margulies, Toshiaki Koike-Akino, Ye Wang, Deniz Erdogmus
MERL Contacts: Toshiaki Koike-Akino; Ye Wang
Research Areas: Artificial Intelligence, Signal Processing, Human-Computer InteractionBrief- The MERL Signal Processing group achieved first place in the cross-subject transfer learning task and fourth place overall in the NeurIPS 2021 BEETL AI Challenge for EEG Transfer Learning. The team included Niklas Smedemark-Margulies (intern from Northeastern University), Toshiaki Koike-Akino, Ye Wang, and Prof. Deniz Erdogmus (Northeastern University). The challenge addresses two types of transfer learning tasks for EEG Biosignals: a homogeneous transfer learning task for cross-subject domain adaptation; and a heterogeneous transfer learning task for cross-data domain adaptation. There were 110+ registered teams in this competition, MERL ranked 1st in the homogeneous transfer learning task, 7th place in the heterogeneous transfer learning task, and 4th place for the combined overall score. For the homogeneous transfer learning task, MERL developed a new pre-shot learning framework based on feature disentanglement techniques for robustness against inter-subject variation to enable calibration-free brain-computer interfaces (BCI). MERL is invited to present our pre-shot learning technique at the NeurIPS 2021 workshop.
MERL Publications
- , "High-Accuracy User Identification Using EEG Biometrics", International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), DOI: 10.1109/EMBC.2016.7590835, August 2016, pp. 854-858.BibTeX TR2016-105 PDF Presentation
- @inproceedings{Koike-Akino2016aug,
- author = {Koike-Akino, Toshiaki and Mahajan, Ruhi and Marks, Tim K. and Tuzel, C. Oncel and Wang, Ye and Watanabe, Shinji and Orlik, Philip V.},
- title = {{High-Accuracy User Identification Using EEG Biometrics}},
- booktitle = {International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC)},
- year = 2016,
- pages = {854--858},
- month = aug,
- doi = {10.1109/EMBC.2016.7590835},
- url = {https://www.merl.com/publications/TR2016-105}
- }
- , "Translating sEMG Signals to Continuous Hand Poses using Recurrent Neural Networks", IEEE Conference on Biomedical and Health Informatics (BHI), DOI: 10.1109/BHI.2018.8333395, January 2018.BibTeX TR2018-014 PDF Presentation
- @inproceedings{Quivira2018jan,
- author = {Quivira, Fernando and Koike-Akino, Toshiaki and Wang, Ye and Erdogmus, Deniz},
- title = {{Translating sEMG Signals to Continuous Hand Poses using Recurrent Neural Networks}},
- booktitle = {IEEE Conference on Biomedical and Health Informatics (BHI)},
- year = 2018,
- month = jan,
- doi = {10.1109/BHI.2018.8333395},
- url = {https://www.merl.com/publications/TR2018-014}
- }
- , "Transfer Learning in Brain-Computer Interfaces with Adversarial Variational Autoencoders", International IEEE EMBS Conference on Neural Engineering, DOI: 10.1109/NER.2019.8716897, March 2019.BibTeX TR2018-184 PDF
- @inproceedings{Ozdenizci2019mar,
- author = {Ozdenizci, Ozan and Wang, Ye and Koike-Akino, Toshiaki and Erdogmus, Deniz},
- title = {{Transfer Learning in Brain-Computer Interfaces with Adversarial Variational Autoencoders}},
- booktitle = {International IEEE EMBS Conference on Neural Engineering},
- year = 2019,
- month = mar,
- doi = {10.1109/NER.2019.8716897},
- url = {https://www.merl.com/publications/TR2018-184}
- }
- , "Spatial Component-wise Convolutional Network (SCCNet) for Motor-Imagery EEG Classification", International IEEE EMBS Conference on Neural Engineering, DOI: 10.1109/NER.2019.8716937, March 2019.BibTeX TR2018-210 PDF
- @inproceedings{Wei2019mar,
- author = {Wei, Chun-Shu and Koike-Akino, Toshiaki and Wang, Ye},
- title = {{Spatial Component-wise Convolutional Network (SCCNet) for Motor-Imagery EEG Classification}},
- booktitle = {International IEEE EMBS Conference on Neural Engineering},
- year = 2019,
- month = mar,
- doi = {10.1109/NER.2019.8716937},
- url = {https://www.merl.com/publications/TR2018-210}
- }
- , "Adversarial Deep Learning in EEG Biometrics", IEEE Signal Processing Letters, DOI: 10.1109/LSP.2019.2906826, Vol. 26, No. 5, pp. 710-714, March 2019.BibTeX TR2019-017 PDF
- @article{Ozdenizci2019mar2,
- author = {Ozdenizci, Ozan and Wang, Ye and Koike-Akino, Toshiaki and Erdogmus, Deniz},
- title = {{Adversarial Deep Learning in EEG Biometrics}},
- journal = {IEEE Signal Processing Letters},
- year = 2019,
- volume = 26,
- number = 5,
- pages = {710--714},
- month = mar,
- doi = {10.1109/LSP.2019.2906826},
- url = {https://www.merl.com/publications/TR2019-017}
- }
- , "Learning Invariant Representations from EEG via Adversarial Inference", IEEE Access, DOI: 10.1109/ACCESS.2020.2971600, Vol. 8, pp. 27074-27085, April 2020.BibTeX TR2020-049 PDF
- @article{Ozdenizci2020apr,
- author = {Ozdenizci, Ozan and Wang, Ye and Koike-Akino, Toshiaki and Erdogmus, Deniz},
- title = {{Learning Invariant Representations from EEG via Adversarial Inference}},
- journal = {IEEE Access},
- year = 2020,
- volume = 8,
- pages = {27074--27085},
- month = apr,
- doi = {10.1109/ACCESS.2020.2971600},
- issn = {2169-3536},
- url = {https://www.merl.com/publications/TR2020-049}
- }
- , "Disentangled Adversarial Transfer Learning for Physiological Biosignals", International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), DOI: 10.1109/EMBC44109.2020.9175233, July 2020.BibTeX TR2020-109 PDF Video Presentation
- @inproceedings{Han2020jul,
- author = {Han, Mo and Ozdenizci, Ozan and Wang, Ye and Koike-Akino, Toshiaki and Erdogmus, Deniz},
- title = {{Disentangled Adversarial Transfer Learning for Physiological Biosignals}},
- booktitle = {International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC)},
- year = 2020,
- month = jul,
- publisher = {IEEE},
- doi = {10.1109/EMBC44109.2020.9175233},
- issn = {1558-4615},
- isbn = {978-1-7281-1990-8},
- url = {https://www.merl.com/publications/TR2020-109}
- }
- , "Disentangled Adversarial Autoencoder for Subject-Invariant Physiological Feature Extraction", IEEE Signal Processing Letters, DOI: 10.1109/LSP.2020.3020215, Vol. 27, pp. 1565-1569, September 2020.BibTeX TR2020-128 PDF
- @article{Han2020sep,
- author = {Han, Mo and Ozdenizci, Ozan and Wang, Ye and Koike-Akino, Toshiaki and Erdogmus, Deniz},
- title = {{Disentangled Adversarial Autoencoder for Subject-Invariant Physiological Feature Extraction}},
- journal = {IEEE Signal Processing Letters},
- year = 2020,
- volume = 27,
- pages = {1565--1569},
- month = sep,
- doi = {10.1109/LSP.2020.3020215},
- issn = {1558-2361},
- url = {https://www.merl.com/publications/TR2020-128}
- }
- , "Universal Physiological Representation Learning with Soft-Disentangled Rateless Autoencoders", IEEE Journal of Biomedical and Health Informatics, DOI: 10.1109/JBHI.2021.3062335, Vol. 25, No. 8, pp. 2928-2937, April 2021.BibTeX TR2021-027 PDF
- @article{Han2021apr,
- author = {Han, Mo and Ozdenizci, Ozan and Koike-Akino, Toshiaki and Wang, Ye and Erdogmus, Deniz},
- title = {{Universal Physiological Representation Learning with Soft-Disentangled Rateless Autoencoders}},
- journal = {IEEE Journal of Biomedical and Health Informatics},
- year = 2021,
- volume = 25,
- number = 8,
- pages = {2928--2937},
- month = apr,
- doi = {10.1109/JBHI.2021.3062335},
- issn = {2168-2208},
- url = {https://www.merl.com/publications/TR2021-027}
- }
- , "AutoBayes: Automated Bayesian Graph Exploration for Nuisance-Robust Inference", IEEE Access, DOI: 10.1109/ACCESS.2021.3064530, Vol. 9, pp. 39955-39972, March 2021.BibTeX TR2021-016 PDF Presentation
- @article{Demir2021mar,
- author = {Demir, Andac and Koike-Akino, Toshiaki and Wang, Ye and Erdogmus, Deniz},
- title = {{AutoBayes: Automated Bayesian Graph Exploration for Nuisance-Robust Inference}},
- journal = {IEEE Access},
- year = 2021,
- volume = 9,
- pages = {39955--39972},
- month = mar,
- doi = {10.1109/ACCESS.2021.3064530},
- issn = {2169-3536},
- url = {https://www.merl.com/publications/TR2021-016}
- }
- , "Stochastic Bottleneck: Rateless Auto-Encoder for Flexible Dimensionality Reduction", IEEE International Symposium on Information Theory (ISIT), DOI: 10.1109/ISIT44484.2020.9174523, June 2020.BibTeX TR2020-075 PDF Video Presentation
- @inproceedings{Koike-Akino2020jun,
- author = {Koike-Akino, Toshiaki and Wang, Ye},
- title = {{Stochastic Bottleneck: Rateless Auto-Encoder for Flexible Dimensionality Reduction}},
- booktitle = {IEEE International Symposium on Information Theory (ISIT)},
- year = 2020,
- month = jun,
- publisher = {IEEE},
- doi = {10.1109/ISIT44484.2020.9174523},
- issn = {2157-8117},
- isbn = {978-1-7281-6432-8},
- url = {https://www.merl.com/publications/TR2020-075}
- }
- , "EEG-GNN: Graph Neural Networks for Classification of Electroencephalogram (EEG) Signals", International IEEE EMBS Conference on Neural Engineering, DOI: 10.1109/EMBC46164.2021.9630194, October 2021.BibTeX TR2021-136 PDF Video Presentation
- @inproceedings{Demir2021oct,
- author = {Demir, Andac and Koike-Akino, Toshiaki and Wang, Ye and Erdogmus, Deniz and Haruna, Masaki},
- title = {{EEG-GNN: Graph Neural Networks for Classification of Electroencephalogram (EEG) Signals}},
- booktitle = {International IEEE EMBS Conference on Neural Engineering},
- year = 2021,
- month = oct,
- publisher = {IEEE},
- doi = {10.1109/EMBC46164.2021.9630194},
- issn = {2694-0604},
- isbn = {978-1-7281-1179-7},
- url = {https://www.merl.com/publications/TR2021-136}
- }
- , "Comparison of Three Feedback Modalities for Haptics Sensation in Remote Machine Manipulation", IEEE International Conference on Robotics and Automation (ICRA), DOI: 10.1109/LRA.2021.3070301, May 2021.BibTeX TR2021-063 PDF Video
- @inproceedings{Haruna2021may,
- author = {Haruna, Masaki and Kawaguchi, Noboru and Ogino, Masaki and Koike-Akino, Toshiaki},
- title = {{Comparison of Three Feedback Modalities for Haptics Sensation in Remote Machine Manipulation}},
- booktitle = {IEEE International Conference on Robotics and Automation (ICRA)},
- year = 2021,
- month = may,
- doi = {10.1109/LRA.2021.3070301},
- issn = {2377-3766},
- url = {https://www.merl.com/publications/TR2021-063}
- }